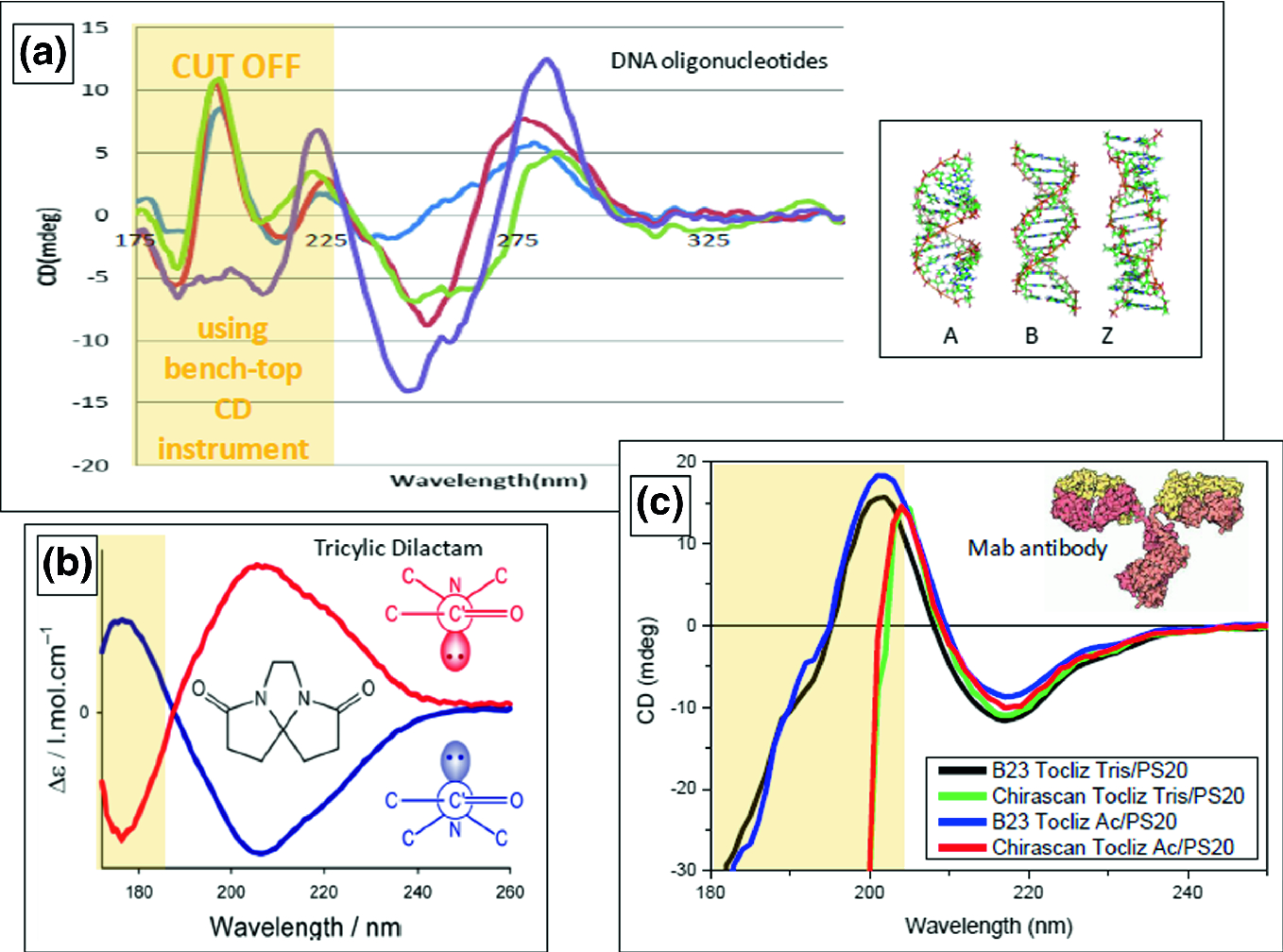
The far-reaching implications of these simple observations are demonstrated through two examples: (i) circular dichroism spectrum deconvolution, and (ii) receptor-ligand affinity determination. However, as IDPs can show extreme variation in amino acid composition and physical features not necessarily covered by our examples, even these techniques should only be used for IDPs following standardization.
By comparison to elemental analysis as the gold standard, we show through the example of four globular proteins and nine IDPs that the ninhydrin assay and the commercial Qubit TM Protein Assay provide reliable data on IDP quantity. Therefore, dependable method(s) have to be worked out/adopted for this task. Routinely used assays for protein quantification, such as the Bradford assay or ultraviolet absorbance at 280 nm, usually seriously misestimate the concentrations of IDPs due to their distinct and variable amino acid composition. Protein quantification is essential in a great variety of biochemical assays, yet the inherent systematic errors associated with the concentration determination of intrinsically disordered proteins (IDPs) using classical methods are hardly appreciated. 3Research Centre for Natural Sciences of the Hungarian Academy of Sciences, Institute of Enzymology, Budapest, Hungary.2Structural Biology Brussels, Vrije Universiteit Brussel, Brussels, Belgium.1VIB-VUB Center for Structural Biology, Vlaams Instituut voor Biotechnologie, Brussels, Belgium.Oemig 1,2, Dominique Maes 2, Kris Pauwels 1,2, Peter Tompa 1,2,3 * and Pierre Lebrun 1,2 Nguyen 1,2 †, Nevena Hristozova 1,2, Mauricio Macossay-Castillo 1,2, Denes Kovacs 1,2, Angela Bekesi 1,2, Jesper S.
